Well done to Helen Horkan for her work contributing to the new v2 publicly available genome for Hydractinia.

Hydractinia is a close relative of sea anemones and corals, it can reproduce by cloning or sexually. In the wild it lives on the shells of hermit crabs as colonies and can be found all across the globe, from Galway Bay to California. Scientists are interested in Hydractinia because it doesn’t age, doesn’t get cancer, has a huge capacity to regenerate lost body parts thanks to its pluripotent stem cells (i-cells). In order to understand Hydractinias interesting abilities scientists use a range of techniques, such as exposing it to various treatments and seeing how it copes with them, manipulating the animals genes and investigating how it produces different cell types, like its stinging cells. In order to carry out these experiments we need to know the details of the animal’s genetic blueprint, or genome. The Uri Frank Lab at University of Galway, in collaboration with the Oleg Simakov Lab at University of Vienna have produced a publicly available genome for Hydractinia, which represents all 15 of the animal’s chromosomes at 66x depth. This genome is a significant step forward for research into Hydractinia and more broadly across the tree of life, as good quality reference genomes for all organisms will help our understanding globally of how humans and other animals evolved and what our Last Universal Common Ancestor may have been.
Fig 1. Hydractinia feeding polyp. piwi1::GFP/bTubulin::mScarlet reporter animal. Somatic cells express red fluorescent protein, pluripotent stem cells (i-cells) express green fluorescent protein.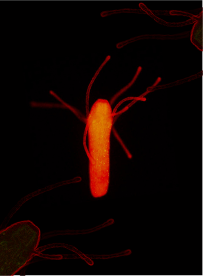
NCBI:
https://www.ncbi.nlm.nih.gov/nuccore/JARBIS000000000.1/
FigShare for all data downloads:

